Since we have already analyzed all these datasets in the target countries section, we see that using the global dataset for all our modeling is the best option for a few reasons:
- it contains data for all the countries we are covering
- it has up to date data that includes the whole lifetime of the pandemic
- the individual countries datasets are not as complete in some cases
- a single dataset is arguably easier to work with compared to many
- the data is already clean and
- we have already confirmed the creadability of the data
Objectives
our main objective is to see which one of our chosen 4 countries have handled the virus in a way that can be generalized to everyone as simple guidelines, the targeted countries are
- United States
- Germany
- Italy
- South Korea
Data Exploration
import pandas as pdimport numpy as npimport matplotlib.pyplot as pltimport tensorflow as tffrom datetime import datetimefrom sklearn.preprocessing import MinMaxScalerfrom keras.preprocessing.sequence import TimeseriesGeneratorfrom keras.models import Sequentialfrom keras.layers import Dense, LSTM, Dropout, Activation, GlobalMaxPooling1D, Bidirectionalfrom keras.optimizers import Adamfrom tensorflow.keras.callbacks import ModelCheckpointfrom statsmodels.tsa.arima_model import ARIMAfrom statsmodels.tsa.api import ExponentialSmoothing, SimpleExpSmoothing, Holt%matplotlib inline# supress annoying warningimport warningsfrom statsmodels.tools.sm_exceptions import ConvergenceWarningwarnings.simplefilter('ignore', ConvergenceWarning)df_confirmed = pd.read_csv("../input/covid-19/time_series_covid19_confirmed_global.csv")df_deaths = pd.read_csv("../input/covid-19/time_series_covid19_deaths_global.csv")df_reco = pd.read_csv("../input/covid-19/time_series_covid19_recovered_global.csv")after reading in our dataset lets take a look at it by showing the first few countries for confirmed case, deaths, and recoveries
df_confirmed.head()| Province/State | Country/Region | Lat | Long | 1/22/20 | 1/23/20 | 1/24/20 | 1/25/20 | 1/26/20 | 1/27/20 | ... | 10/22/20 | 10/23/20 | 10/24/20 | 10/25/20 | 10/26/20 | 10/27/20 | 10/28/20 | 10/29/20 | 10/30/20 | 10/31/20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | Afghanistan | 33.93911 | 67.709953 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 40626 | 40687 | 40768 | 40833 | 40937 | 41032 | 41145 | 41268 | 41334 | 41425 |
| 1 | NaN | Albania | 41.15330 | 20.168300 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 18250 | 18556 | 18858 | 19157 | 19445 | 19729 | 20040 | 20315 | 20634 | 20875 |
| 2 | NaN | Algeria | 28.03390 | 1.659600 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 55357 | 55630 | 55880 | 56143 | 56419 | 56706 | 57026 | 57332 | 57651 | 57942 |
| 3 | NaN | Andorra | 42.50630 | 1.521800 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 3811 | 4038 | 4038 | 4038 | 4325 | 4410 | 4517 | 4567 | 4665 | 4756 |
| 4 | NaN | Angola | -11.20270 | 17.873900 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 8582 | 8829 | 9026 | 9381 | 9644 | 9871 | 10074 | 10269 | 10558 | 10805 |
5 rows × 288 columns
df_deaths.head()| Province/State | Country/Region | Lat | Long | 1/22/20 | 1/23/20 | 1/24/20 | 1/25/20 | 1/26/20 | 1/27/20 | ... | 10/22/20 | 10/23/20 | 10/24/20 | 10/25/20 | 10/26/20 | 10/27/20 | 10/28/20 | 10/29/20 | 10/30/20 | 10/31/20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | Afghanistan | 33.93911 | 67.709953 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 1505 | 1507 | 1511 | 1514 | 1518 | 1523 | 1529 | 1532 | 1533 | 1536 |
| 1 | NaN | Albania | 41.15330 | 20.168300 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 465 | 469 | 473 | 477 | 480 | 487 | 493 | 499 | 502 | 509 |
| 2 | NaN | Algeria | 28.03390 | 1.659600 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 1888 | 1897 | 1907 | 1914 | 1922 | 1931 | 1941 | 1949 | 1956 | 1964 |
| 3 | NaN | Andorra | 42.50630 | 1.521800 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 63 | 69 | 69 | 69 | 72 | 72 | 72 | 73 | 75 | 75 |
| 4 | NaN | Angola | -11.20270 | 17.873900 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 260 | 265 | 267 | 268 | 270 | 271 | 275 | 275 | 279 | 284 |
5 rows × 288 columns
df_reco.head()| Province/State | Country/Region | Lat | Long | 1/22/20 | 1/23/20 | 1/24/20 | 1/25/20 | 1/26/20 | 1/27/20 | ... | 10/22/20 | 10/23/20 | 10/24/20 | 10/25/20 | 10/26/20 | 10/27/20 | 10/28/20 | 10/29/20 | 10/30/20 | 10/31/20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | Afghanistan | 33.93911 | 67.709953 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 33831 | 34010 | 34023 | 34129 | 34150 | 34217 | 34237 | 34239 | 34258 | 34321 |
| 1 | NaN | Albania | 41.15330 | 20.168300 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 10395 | 10466 | 10548 | 10654 | 10705 | 10808 | 10893 | 11007 | 11097 | 11189 |
| 2 | NaN | Algeria | 28.03390 | 1.659600 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 38618 | 38788 | 38932 | 39095 | 39273 | 39444 | 39635 | 39635 | 40014 | 40201 |
| 3 | NaN | Andorra | 42.50630 | 1.521800 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 2470 | 2729 | 2729 | 2729 | 2957 | 3029 | 3144 | 3260 | 3377 | 3475 |
| 4 | NaN | Angola | -11.20270 | 17.873900 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 3305 | 3384 | 3461 | 3508 | 3530 | 3647 | 3693 | 3736 | 4107 | 4523 |
5 rows × 288 columns
after taking a look at the data as a whole lets now get our target countries each in their own dataframes
us_confirmed = df_confirmed[df_confirmed["Country/Region"] == "US"]us_deaths = df_deaths[df_deaths["Country/Region"] == "US"]us_reco = df_reco[df_reco["Country/Region"] == "US"]germany_confirmed = df_confirmed[df_confirmed["Country/Region"] == "Germany"]germany_deaths = df_deaths[df_deaths["Country/Region"] == "Germany"]germany_reco = df_reco[df_reco["Country/Region"] == "Germany"]italy_confirmed = df_confirmed[df_confirmed["Country/Region"] == "Italy"]italy_deaths = df_deaths[df_deaths["Country/Region"] == "Italy"]italy_reco = df_reco[df_reco["Country/Region"] == "Italy"]sk_confirmed = df_confirmed[df_confirmed["Country/Region"] == "Korea, South"]sk_deaths = df_deaths[df_deaths["Country/Region"] == "Korea, South"]sk_reco = df_reco[df_reco["Country/Region"] == "Korea, South"]us_reco| Province/State | Country/Region | Lat | Long | 1/22/20 | 1/23/20 | 1/24/20 | 1/25/20 | 1/26/20 | 1/27/20 | ... | 10/22/20 | 10/23/20 | 10/24/20 | 10/25/20 | 10/26/20 | 10/27/20 | 10/28/20 | 10/29/20 | 10/30/20 | 10/31/20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 231 | NaN | US | 40.0 | -100.0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 3353056 | 3375427 | 3406656 | 3422878 | 3460455 | 3487666 | 3518140 | 3554336 | 3578452 | 3612478 |
1 rows × 288 columns
with the current data structure shown above we cant do much so lets first convert it to a form that can used to make graphs or train a model
## structuring timeseries datadef confirmed_timeseries(df): df_series = pd.DataFrame(df[df.columns[4:]].sum(),columns=["confirmed"]) df_series.index = pd.to_datetime(df_series.index,format = '%m/%d/%y') return df_seriesdef deaths_timeseries(df): df_series = pd.DataFrame(df[df.columns[4:]].sum(),columns=["deaths"]) df_series.index = pd.to_datetime(df_series.index,format = '%m/%d/%y') return df_seriesdef reco_timeseries(df): # no index to timeseries conversion needed (all is joined later) df_series = pd.DataFrame(df[df.columns[4:]].sum(),columns=["recovered"]) return df_seriesus_con_series = confirmed_timeseries(us_confirmed)us_dea_series = deaths_timeseries(us_deaths)us_reco_series = reco_timeseries(us_reco)germany_con_series = confirmed_timeseries(germany_confirmed)germany_dea_series = deaths_timeseries(germany_deaths)germany_reco_series = reco_timeseries(germany_reco)italy_con_series = confirmed_timeseries(italy_confirmed)italy_dea_series = deaths_timeseries(italy_deaths)italy_reco_series = reco_timeseries(italy_reco)sk_con_series = confirmed_timeseries(sk_confirmed)sk_dea_series = deaths_timeseries(sk_deaths)sk_reco_series = reco_timeseries(sk_reco)# join all data frames for each county (makes it easier to graph and compare)us_df = us_con_series.join(us_dea_series, how = "inner")us_df = us_df.join(us_reco_series, how = "inner")germany_df = germany_con_series.join(germany_dea_series, how = "inner")germany_df = germany_df.join(germany_reco_series, how = "inner")italy_df = italy_con_series.join(italy_dea_series, how = "inner")italy_df = italy_df.join(italy_reco_series, how = "inner")sk_df = sk_con_series.join(sk_dea_series, how = "inner")sk_df = sk_df.join(sk_reco_series, how = "inner")us_df| confirmed | deaths | recovered | |
|---|---|---|---|
| 2020-01-22 | 1 | 0 | 0 |
| 2020-01-23 | 1 | 0 | 0 |
| 2020-01-24 | 2 | 0 | 0 |
| 2020-01-25 | 2 | 0 | 0 |
| 2020-01-26 | 5 | 0 | 0 |
| ... | ... | ... | ... |
| 2020-10-27 | 8778055 | 226696 | 3487666 |
| 2020-10-28 | 8856413 | 227685 | 3518140 |
| 2020-10-29 | 8944934 | 228656 | 3554336 |
| 2020-10-30 | 9044255 | 229686 | 3578452 |
| 2020-10-31 | 9125482 | 230548 | 3612478 |
284 rows × 3 columns
Visual and Descriptive Analysis
data visualization and descriptive analysis for each country
USA
us_df.plot(figsize=(14,7),title="United States confirmed, deaths and recoverd cases")<matplotlib.axes._subplots.AxesSubplot at 0x7f541b66df50>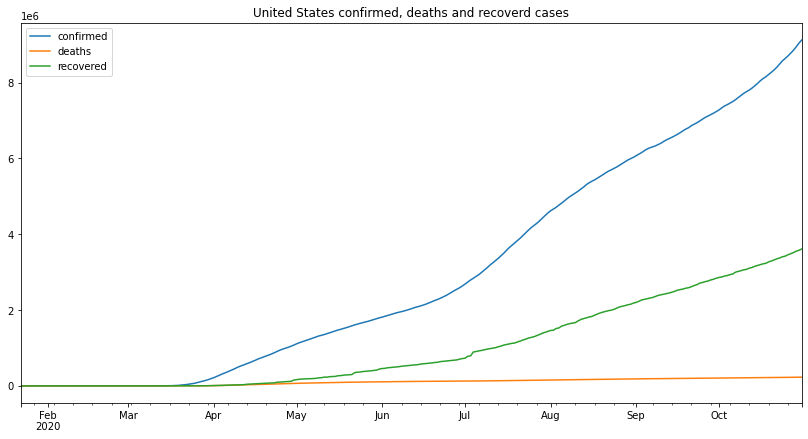
the number of confirmed cases started up slow until around April, it started to go up at a much faster rate and it kept that pace even during quarantine, in July the rate at which the cases are increasing got higher and the cases started increasing faster, this can be attributed to the recent protests and people’s ignorance to the CDC guidelines.
deaths are the only cases that have had a continuously increasing rate, all the way from April the number of deaths is increasing at an increasing rate, until august where the increase rate is slower despite the higher number of cases.
when it comes to the recoveries, the recovery starts at the same time as the confirmed cases with a very unstable increase rate, the highest increase rate is also from around July which is surprising considering the rate of confirmed cases also went up around that time.
us_cases_outcome = (us_df.tail(1)['deaths'] + us_df.tail(1)['recovered'])[0]us_outcome_perc = (us_cases_outcome / us_df.tail(1)['confirmed'] * 100)[0]us_death_perc = (us_df.tail(1)['deaths'] / us_cases_outcome * 100)[0]us_reco_perc = (us_df.tail(1)['recovered'] / us_cases_outcome * 100)[0]us_active = (us_df.tail(1)['confirmed'] - us_cases_outcome)[0]print(f"Number of cases which had an outcome: {us_cases_outcome}")print(f"percentage of cases that had an outcome: {round(us_outcome_perc, 2)}%")print(f"Deaths rate: {round(us_death_perc, 2)}%")print(f"Recovery rate: {round(us_reco_perc, 2)}%")print(f"Currently Active cases: {us_active}")Number of cases which had an outcome: 3843026percentage of cases that had an outcome: 42.11%Deaths rate: 6.0%Recovery rate: 94.0%Currently Active cases: 5282456the percentage of cases that had an outcome is just 38.06% of the total cases, which is very low, the other 61.4 of the cases which are not accounted for have probably not been released officially by the government, however, the recovery rate is high at 91.79% while the death rate is at 8.21%
number of currently active cases is still very high, and it’s going up if the current increase rates are to be quoted.
Modeling
for modeling and predicting the number of cases in the upcoming days the following types of models will be implemented:
- Bidrectional Long Short Term Memory (BiLSTM)
LSTMs’ are known and widely used in time sensitive data where a variable is increaing with time depending on the values from prior days.
- Autoregressive Integrated Moving Average (ARIMA)
models the next step in the sequence as a linear function of the observations and resiudal errors at prior time steps.
- Holt’s Exponential Smoothing (HES)
also referred to as holt’s linear trend model or double exponential smoothing, models the next time step as an exponentially weighted linear function of observations at prior time step taking into account trends (the only difference from SES)
each country will have a total number of 3 models and the results will be compared accordingly.
our data is in a daily format and we want to predict n days at a time so we will take out the last n days and use them to test and predict outcomes it 2 weeks time.
n_input = 10 # number of stepsn_features = 1 # number of y# prepare required input datadef prepare_data(df): # drop rows with zeros df = df[(df.T != 0).any()] num_days = len(df) - n_input train = df.iloc[:num_days] test = df.iloc[num_days:] # normalize the data according to largest value scaler = MinMaxScaler() scaler.fit(train) # find max value scaled_train = scaler.transform(train) # divide every point by max value scaled_test = scaler.transform(test) # feed in batches [t1,t2,t3] --> t4 generator = TimeseriesGenerator(scaled_train,scaled_train,length = n_input,batch_size = 1) validation_set = np.append(scaled_train[55],scaled_test) # random tbh validation_set = validation_set.reshape(n_input + 1,1) validation_gen = TimeseriesGenerator(validation_set,validation_set,length = n_input,batch_size = 1) return scaler, train, test, scaled_train, scaled_test, generator, validation_genBuilding the models
# create, train and return LSTM modeldef train_lstm_model(): model = Sequential() model.add(Bidirectional(LSTM(84, recurrent_dropout = 0, unroll = False, return_sequences = True, use_bias = True, input_shape = (n_input,n_features)))) model.add(LSTM(84, recurrent_dropout = 0.1, use_bias = True, return_sequences = True,)) model.add(GlobalMaxPooling1D()) model.add(Dense(84, activation = "relu")) model.add(Dense(units = 1)) # compile model model.compile(loss = 'mae', optimizer = Adam(1e-5)) # finally train the model using generators model.fit_generator(generator,validation_data = validation_gen, epochs = 100, steps_per_epoch = round(len(train) / n_input), verbose = 0) return model# predict, rescale and append needed columns to final data framedef lstm_predict(model): # holding predictions test_prediction = [] # last n points from training set first_eval_batch = scaled_train[-n_input:] current_batch = first_eval_batch.reshape(1,n_input,n_features) # predict first x days from testing data for i in range(len(test) + n_input): current_pred = model.predict(current_batch)[0] test_prediction.append(current_pred) current_batch = np.append(current_batch[:,1:,:],[[current_pred]],axis=1) # inverse scaled data true_prediction = scaler.inverse_transform(test_prediction) MAPE, accuracy, sum_errs, interval, stdev, df_forecast = gen_metrics(true_prediction) return MAPE, accuracy, sum_errs, interval, stdev, df_forecast# plotting model lossesdef plot_lstm_losses(model): pd.DataFrame(model.history.history).plot(figsize = (14,7), title = "loss vs epochs curve")'''incrementally trained ARIMA: - train with original train data - predict the next value - appened the prediction value to the training data - repeat training and appending for n times (days in this case) this incremental technique significantly improves the accuracy by always using all data up to previous day for predeicting next value unlike predecting multiple values at the same time which is not incremeital. PARAMETERS: p: autoregressive(AR) order d: order of differencing q: moving average(MA) order'''def arima_predict(p: int, d: int, q: int): values = [x for x in train.values] predictions = [] for t in range(len(test) + n_input): # the number of testing days + the future days to predict model = ARIMA(values, order = (p,d,q)) model_fit = model.fit() fcast = model_fit.forecast() predictions.append(fcast[0][0]) values.append(fcast[0]) MAPE, accuracy, sum_errs, interval, stdev, df_forecast = gen_metrics(predictions) return MAPE, accuracy, sum_errs, interval, stdev, df_forecast'''incremental Holt's (Method) Exponential Smoothing - trained the same way as above arima'''def hes_predict(): values = [x for x in train.values] predictions = [] for t in range(len(test) + n_input): # the number of testing days + the future days to predict model = Holt(values) model_fit = model.fit() fcast = model_fit.predict() predictions.append(fcast[0]) values.append(fcast[0]) MAPE, accuracy, sum_errs, interval, stdev, df_forecast = gen_metrics(predictions) return MAPE, accuracy, sum_errs, interval, stdev, df_forecast# generate a dataframe with given rangedef get_range_df(start: str, end: str, df): target_df = df.loc[pd.to_datetime(start, format='%Y-%m-%d'):pd.to_datetime(end, format='%Y-%m-%d')] return target_df# fill na values in a range predicted data frame with actual values from the original dataframedef pad_range_df(df, original_df): df['confirmed'] = df.confirmed.fillna(original_df['confirmed']) # fill confirmed Na # fill daily na daily_act = [] daily_df = pd.DataFrame(columns = ["daily"], index = df[n_input:].index) for num in range(n_input - 1, (n_input * 2) - 1): daily_act.append(df["confirmed"].iloc[num + 1] - df["confirmed"].iloc[num]) daily_df['daily'] = daily_act df['daily'] = df.daily.fillna(daily_df['daily']) return df# generate metrics and final dfdef gen_metrics(pred): # create time series time_series_array = test.index for k in range(0, n_input): time_series_array = time_series_array.append(time_series_array[-1:] + pd.DateOffset(1)) # create time series data frame df_forecast = pd.DataFrame(columns = ["confirmed","confirmed_predicted"],index = time_series_array) # append confirmed and predicted confirmed df_forecast.loc[:,"confirmed_predicted"] = pred df_forecast.loc[:,"confirmed"] = test["confirmed"] # create and append daily cases (for both actual and predicted) daily_act = [] daily_pred = [] #actual daily_act.append(abs(df_forecast["confirmed"].iloc[1] - train["confirmed"].iloc[-1])) for num in range((n_input * 2) - 1): daily_act.append(df_forecast["confirmed"].iloc[num + 1] - df_forecast["confirmed"].iloc[num]) # predicted daily_pred.append(df_forecast["confirmed_predicted"].iloc[1] - train["confirmed"].iloc[-1]) for num in range((n_input * 2) - 1): daily_pred.append(df_forecast["confirmed_predicted"].iloc[num + 1] - df_forecast["confirmed_predicted"].iloc[num]) df_forecast["daily"] = daily_act df_forecast["daily_predicted"] = daily_pred # calculate mean absolute percentage error MAPE = np.mean(np.abs(np.array(df_forecast["confirmed"][:n_input]) - np.array(df_forecast["confirmed_predicted"][:n_input])) / np.array(df_forecast["confirmed"][:n_input])) accuracy = round((1 - MAPE) * 100, 2) # the error rate sum_errs = np.sum((np.array(df_forecast["confirmed"][:n_input]) - np.array(df_forecast["confirmed_predicted"][:n_input])) ** 2) # error standard deviation stdev = np.sqrt(1 / (n_input - 2) * sum_errs) # calculate prediction interval interval = 1.96 * stdev # append the min and max cases to final df df_forecast["confirm_min"] = df_forecast["confirmed_predicted"] - interval df_forecast["confirm_max"] = df_forecast["confirmed_predicted"] + interval # round all df values to 0 decimal points df_forecast = df_forecast.round() return MAPE, accuracy, sum_errs, interval, stdev, df_forecast# print metrics for given countydef print_metrics(mape, accuracy, errs, interval, std, model_type): m_str = "LSTM" if model_type == 0 else "ARIMA" if model_type == 1 else "HES" print(f"{m_str} MAPE: {round(mape * 100, 2)}%") print(f"{m_str} accuracy: {accuracy}%") print(f"{m_str} sum of errors: {round(errs)}") print(f"{m_str} prediction interval: {round(interval)}") print(f"{m_str} standard deviation: {std}")# for plotting the range of predicetionsdef plot_results(df, country, algo): fig, (ax1, ax2) = plt.subplots(2, figsize = (14,20)) ax1.set_title(f"{country} {algo} confirmed predictions") ax1.plot(df.index,df["confirmed"], label = "confirmed") ax1.plot(df.index,df["confirmed_predicted"], label = "confirmed_predicted") ax1.fill_between(df.index,df["confirm_min"], df["confirm_max"], color = "indigo",alpha = 0.09,label = "Confidence Interval") ax1.legend(loc = 2) ax2.set_title(f"{country} {algo} confirmed daily predictions") ax2.plot(df.index, df["daily"], label = "daily") ax2.plot(df.index, df["daily_predicted"], label = "daily_predicted") ax2.legend() import matplotlib.dates as mdates ax1.xaxis.set_major_formatter(mdates.DateFormatter('%b %-d')) ax2.xaxis.set_major_formatter(mdates.DateFormatter('%b %-d')) fig.show()USA Predictions
# prepare the datascaler, train, test, scaled_train, scaled_test, generator, validation_gen = prepare_data(us_con_series)# train lstm modelus_lstm_model = train_lstm_model()# plot lstm lossesplot_lstm_losses(us_lstm_model)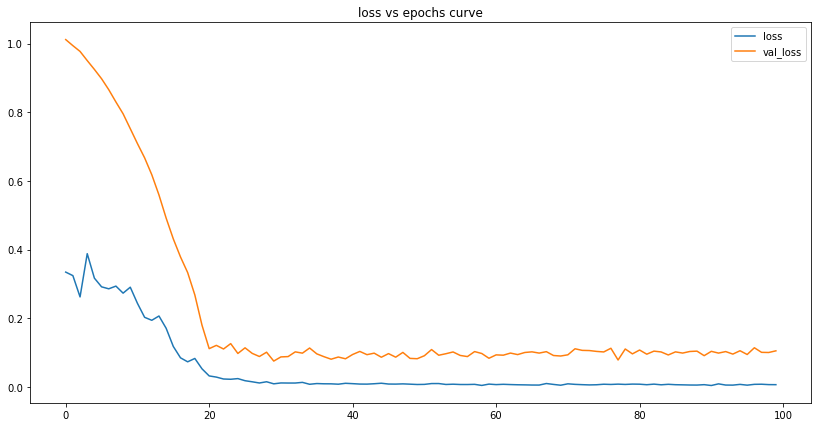
# Long short memory methodus_mape, us_accuracy, us_errs, us_interval, us_std, us_lstm_df = lstm_predict(us_lstm_model)print_metrics(us_mape, us_accuracy, us_errs, us_interval, us_std, 0)us_lstm_dfLSTM MAPE: 4.82%LSTM accuracy: 95.18%LSTM sum of errors: 2058053740601.0LSTM prediction interval: 994121.0LSTM standard deviation: 507204.8083122943| confirmed | confirmed_predicted | daily | daily_predicted | confirm_min | confirm_max | |
|---|---|---|---|---|---|---|
| 2020-10-22 | 8409341.0 | 8205034.0 | 155448.0 | -94666.0 | 7210912.0 | 9199155.0 |
| 2020-10-23 | 8493088.0 | 8242974.0 | 83747.0 | 37941.0 | 7248853.0 | 9237096.0 |
| 2020-10-24 | 8576818.0 | 8276551.0 | 83730.0 | 33577.0 | 7282430.0 | 9270672.0 |
| 2020-10-25 | 8637625.0 | 8306049.0 | 60807.0 | 29498.0 | 7311928.0 | 9300171.0 |
| 2020-10-26 | 8704423.0 | 8331693.0 | 66798.0 | 25644.0 | 7337572.0 | 9325815.0 |
| 2020-10-27 | 8778055.0 | 8354006.0 | 73632.0 | 22313.0 | 7359885.0 | 9348127.0 |
| 2020-10-28 | 8856413.0 | 8373629.0 | 78358.0 | 19623.0 | 7379508.0 | 9367750.0 |
| 2020-10-29 | 8944934.0 | 8392144.0 | 88521.0 | 18515.0 | 7398022.0 | 9386265.0 |
| 2020-10-30 | 9044255.0 | 8408804.0 | 99321.0 | 16660.0 | 7414683.0 | 9402925.0 |
| 2020-10-31 | 9125482.0 | 8423733.0 | 81227.0 | 14929.0 | 7429611.0 | 9417854.0 |
| 2020-11-01 | NaN | 8435789.0 | NaN | 12056.0 | 7441668.0 | 9429910.0 |
| 2020-11-02 | NaN | 8456020.0 | NaN | 20231.0 | 7461899.0 | 9450142.0 |
| 2020-11-03 | NaN | 8474728.0 | NaN | 18708.0 | 7480607.0 | 9468849.0 |
| 2020-11-04 | NaN | 8492286.0 | NaN | 17558.0 | 7498164.0 | 9486407.0 |
| 2020-11-05 | NaN | 8509005.0 | NaN | 16720.0 | 7514884.0 | 9503127.0 |
| 2020-11-06 | NaN | 8525149.0 | NaN | 16143.0 | 7531027.0 | 9519270.0 |
| 2020-11-07 | NaN | 8540892.0 | NaN | 15744.0 | 7546771.0 | 9535014.0 |
| 2020-11-08 | NaN | 8556397.0 | NaN | 15504.0 | 7562275.0 | 9550518.0 |
| 2020-11-09 | NaN | 8571668.0 | NaN | 15272.0 | 7577547.0 | 9565790.0 |
| 2020-11-10 | NaN | 8586776.0 | NaN | 15108.0 | 7592655.0 | 9580897.0 |
plot_results(us_lstm_df, "USA", "LSTM")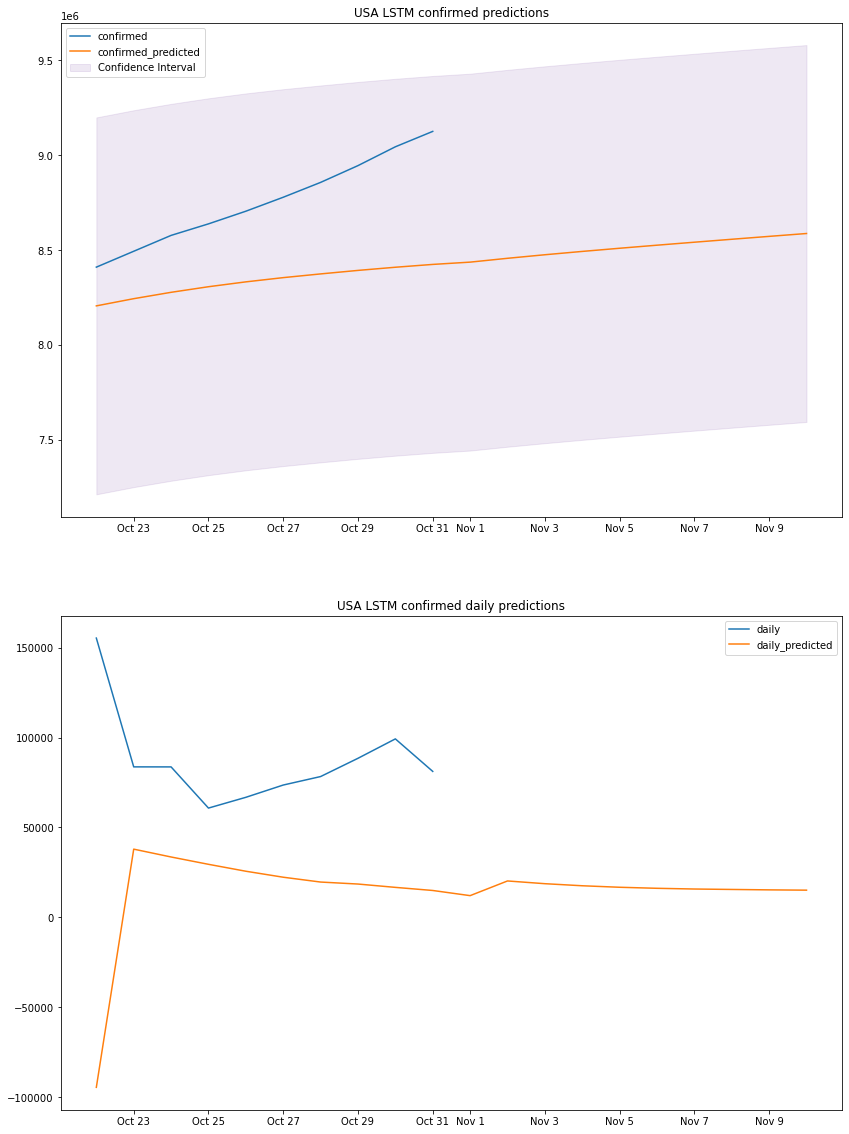
# Auto Regressive Integrated Moving Averageus_mape, us_accuracy, us_errs, us_interval, us_std, us_arima_df = arima_predict(8, 1, 1)print_metrics(us_mape, us_accuracy, us_errs, us_interval, us_std, 1)us_arima_dfARIMA MAPE: 0.72%ARIMA accuracy: 99.28%ARIMA sum of errors: 58884624871.0ARIMA prediction interval: 168156.0ARIMA standard deviation: 85793.81160044324| confirmed | confirmed_predicted | daily | daily_predicted | confirm_min | confirm_max | |
|---|---|---|---|---|---|---|
| 2020-10-22 | 8409341.0 | 8406811.0 | 155448.0 | 138884.0 | 8238655.0 | 8574967.0 |
| 2020-10-23 | 8493088.0 | 8476524.0 | 83747.0 | 69713.0 | 8308368.0 | 8644680.0 |
| 2020-10-24 | 8576818.0 | 8536828.0 | 83730.0 | 60303.0 | 8368672.0 | 8704983.0 |
| 2020-10-25 | 8637625.0 | 8593830.0 | 60807.0 | 57002.0 | 8425674.0 | 8761986.0 |
| 2020-10-26 | 8704423.0 | 8654269.0 | 66798.0 | 60439.0 | 8486113.0 | 8822425.0 |
| 2020-10-27 | 8778055.0 | 8716789.0 | 73632.0 | 62520.0 | 8548633.0 | 8884945.0 |
| 2020-10-28 | 8856413.0 | 8782879.0 | 78358.0 | 66089.0 | 8614723.0 | 8951034.0 |
| 2020-10-29 | 8944934.0 | 8853301.0 | 88521.0 | 70422.0 | 8685145.0 | 9021457.0 |
| 2020-10-30 | 9044255.0 | 8921778.0 | 99321.0 | 68477.0 | 8753622.0 | 9089933.0 |
| 2020-10-31 | 9125482.0 | 8984015.0 | 81227.0 | 62237.0 | 8815859.0 | 9152170.0 |
| 2020-11-01 | NaN | 9043905.0 | NaN | 59891.0 | 8875750.0 | 9212061.0 |
| 2020-11-02 | NaN | 9104850.0 | NaN | 60944.0 | 8936694.0 | 9273006.0 |
| 2020-11-03 | NaN | 9167791.0 | NaN | 62941.0 | 8999635.0 | 9335947.0 |
| 2020-11-04 | NaN | 9234322.0 | NaN | 66531.0 | 9066166.0 | 9402478.0 |
| 2020-11-05 | NaN | 9303277.0 | NaN | 68955.0 | 9135121.0 | 9471433.0 |
| 2020-11-06 | NaN | 9369836.0 | NaN | 66560.0 | 9201681.0 | 9537992.0 |
| 2020-11-07 | NaN | 9431964.0 | NaN | 62128.0 | 9263808.0 | 9600120.0 |
| 2020-11-08 | NaN | 9491890.0 | NaN | 59926.0 | 9323734.0 | 9660046.0 |
| 2020-11-09 | NaN | 9551959.0 | NaN | 60069.0 | 9383803.0 | 9720114.0 |
| 2020-11-10 | NaN | 9613919.0 | NaN | 61961.0 | 9445763.0 | 9782075.0 |
plot_results(us_arima_df, "USA", "incremental ARIMA")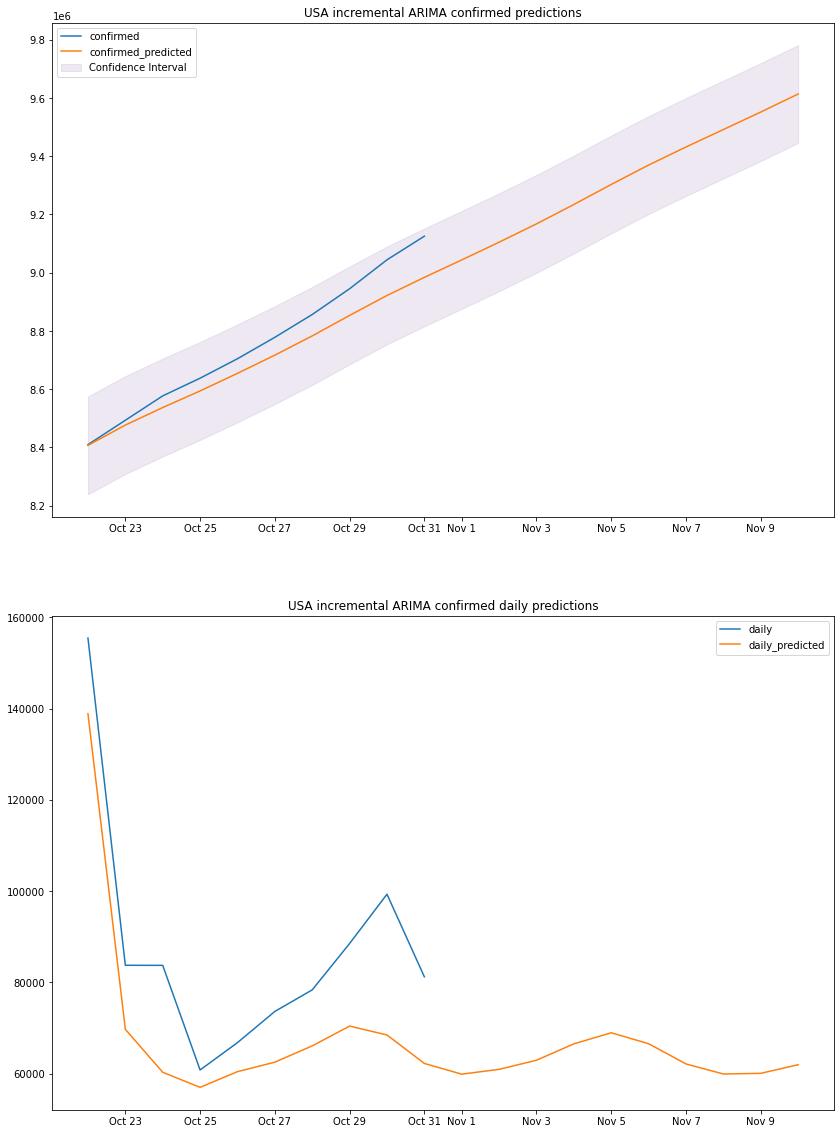
# Holts Exponential Smoothingus_mape, us_accuracy, us_errs, us_interval, us_std, us_hes_df = hes_predict()print_metrics(us_mape, us_accuracy, us_errs, us_interval, us_std, 2)us_hes_dfHES MAPE: 0.83%HES accuracy: 99.17%HES sum of errors: 75839516016.0HES prediction interval: 190835.0HES standard deviation: 97364.98088117794| confirmed | confirmed_predicted | daily | daily_predicted | confirm_min | confirm_max | |
|---|---|---|---|---|---|---|
| 2020-10-22 | 8409341.0 | 8400415.0 | 155448.0 | 125551.0 | 8209580.0 | 8591251.0 |
| 2020-10-23 | 8493088.0 | 8463191.0 | 83747.0 | 62776.0 | 8272356.0 | 8654027.0 |
| 2020-10-24 | 8576818.0 | 8525967.0 | 83730.0 | 62776.0 | 8335132.0 | 8716802.0 |
| 2020-10-25 | 8637625.0 | 8588743.0 | 60807.0 | 62776.0 | 8397907.0 | 8779578.0 |
| 2020-10-26 | 8704423.0 | 8651518.0 | 66798.0 | 62776.0 | 8460683.0 | 8842354.0 |
| 2020-10-27 | 8778055.0 | 8714294.0 | 73632.0 | 62776.0 | 8523459.0 | 8905129.0 |
| 2020-10-28 | 8856413.0 | 8777070.0 | 78358.0 | 62776.0 | 8586234.0 | 8967905.0 |
| 2020-10-29 | 8944934.0 | 8839845.0 | 88521.0 | 62776.0 | 8649010.0 | 9030681.0 |
| 2020-10-30 | 9044255.0 | 8902621.0 | 99321.0 | 62776.0 | 8711786.0 | 9093457.0 |
| 2020-10-31 | 9125482.0 | 8965397.0 | 81227.0 | 62776.0 | 8774562.0 | 9156232.0 |
| 2020-11-01 | NaN | 9028173.0 | NaN | 62776.0 | 8837337.0 | 9219008.0 |
| 2020-11-02 | NaN | 9090948.0 | NaN | 62776.0 | 8900113.0 | 9281784.0 |
| 2020-11-03 | NaN | 9153724.0 | NaN | 62776.0 | 8962889.0 | 9344559.0 |
| 2020-11-04 | NaN | 9216500.0 | NaN | 62776.0 | 9025664.0 | 9407335.0 |
| 2020-11-05 | NaN | 9279275.0 | NaN | 62776.0 | 9088440.0 | 9470111.0 |
| 2020-11-06 | NaN | 9342051.0 | NaN | 62776.0 | 9151216.0 | 9532887.0 |
| 2020-11-07 | NaN | 9404827.0 | NaN | 62776.0 | 9213992.0 | 9595662.0 |
| 2020-11-08 | NaN | 9467603.0 | NaN | 62776.0 | 9276767.0 | 9658438.0 |
| 2020-11-09 | NaN | 9530378.0 | NaN | 62776.0 | 9339543.0 | 9721214.0 |
| 2020-11-10 | NaN | 9593154.0 | NaN | 62776.0 | 9402319.0 | 9783989.0 |
plot_results(us_hes_df, "USA", "incremental HES")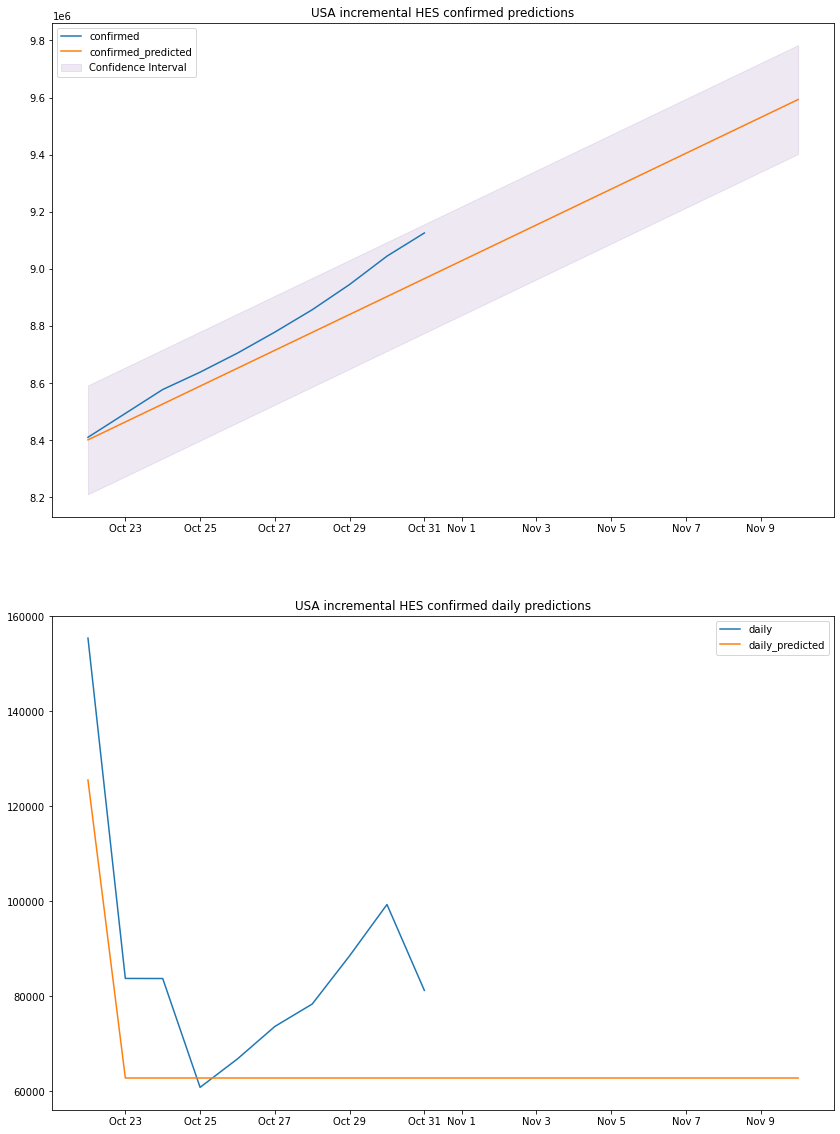
Effectiveness of mandated lockdown
was the US lockdown effective in reducing the cases?
the US started their lockdwon in 2020-03-17 and it was ended by the erupting
protests. tracking the lockdown might be tricky in the US at least because each
state started their lockdown at their own pace and there was no federally
mandated lockdown while some other states never went into lockdowns, taking that
into account we will consider the end of the lockdown to be the end of may which
was the start of the Gorge Floyed protests.
Time frame from 2020-03-17 until 2020-05-31
us_lockdown = get_range_df('2020-03-17', '2020-05-31', us_con_series)fig, ax = plt.subplots(1, figsize = (14,7))ax.plot(us_lockdown.index, us_lockdown['confirmed'], label = 'confirmed')ax.plot(us_lockdown.index, us_lockdown.rolling(7).mean(), label = 'confirmed mean')ax.legend()<matplotlib.legend.Legend at 0x7f53d0630f90>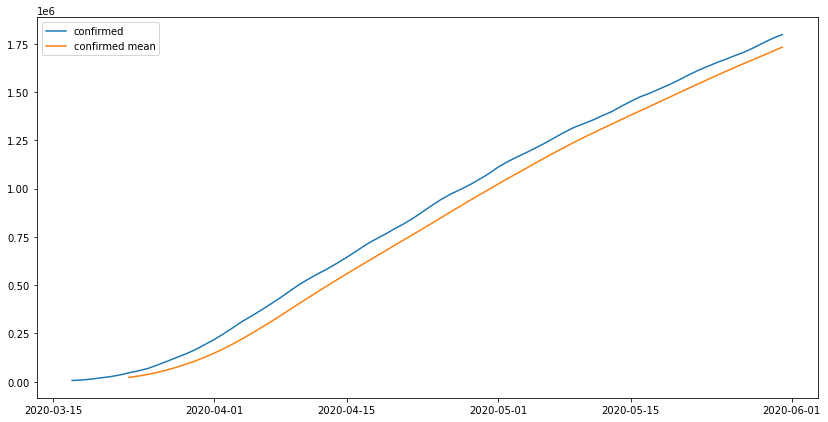
the actual values are above the moving average of each 7 days meaning the lockdown did not work as inteded and the number of cases was still very high when compared to the average of each 7 days, to make sure our previous model predictions are accurate we will use this period of time as a base and train the model on it and do prediction for the days after that which we already have the data on. we will use the ARIMA model becuase the amount of data we have will not train a neural network ideally.
scaler, train, test, scaled_train, scaled_test, generator, validation_gen = prepare_data(us_lockdown)# Auto Regressive Integrated Moving Averageus_mape, us_accuracy, us_errs, us_interval, us_std, us_arima_df = arima_predict(8, 1, 1)print_metrics(us_mape, us_accuracy, us_errs, us_interval, us_std, 1)us_arima_df = pad_range_df(us_arima_df, us_con_series)us_arima_dfARIMA MAPE: 0.55%ARIMA accuracy: 99.45%ARIMA sum of errors: 1187672403.0ARIMA prediction interval: 23881.0ARIMA standard deviation: 12184.377305753513| confirmed | confirmed_predicted | daily | daily_predicted | confirm_min | confirm_max | |
|---|---|---|---|---|---|---|
| 2020-05-22 | 1608604.0 | 1610272.0 | 44636.0 | 47633.0 | 1586391.0 | 1634154.0 |
| 2020-05-23 | 1629802.0 | 1632799.0 | 21198.0 | 22527.0 | 1608918.0 | 1656681.0 |
| 2020-05-24 | 1649916.0 | 1653311.0 | 20114.0 | 20511.0 | 1629429.0 | 1677192.0 |
| 2020-05-25 | 1668235.0 | 1674419.0 | 18319.0 | 21108.0 | 1650537.0 | 1698300.0 |
| 2020-05-26 | 1687761.0 | 1695924.0 | 19526.0 | 21505.0 | 1672042.0 | 1719805.0 |
| 2020-05-27 | 1706351.0 | 1719538.0 | 18590.0 | 23614.0 | 1695657.0 | 1743419.0 |
| 2020-05-28 | 1729299.0 | 1744487.0 | 22948.0 | 24949.0 | 1720606.0 | 1768369.0 |
| 2020-05-29 | 1753651.0 | 1768770.0 | 24352.0 | 24283.0 | 1744889.0 | 1792652.0 |
| 2020-05-30 | 1777495.0 | 1791120.0 | 23844.0 | 22350.0 | 1767239.0 | 1815002.0 |
| 2020-05-31 | 1796670.0 | 1812184.0 | 19175.0 | 21064.0 | 1788303.0 | 1836066.0 |
| 2020-06-01 | 1814034.0 | 1833132.0 | 17364.0 | 20948.0 | 1809251.0 | 1857014.0 |
| 2020-06-02 | 1835408.0 | 1854887.0 | 21374.0 | 21755.0 | 1831006.0 | 1878768.0 |
| 2020-06-03 | 1855386.0 | 1878310.0 | 19978.0 | 23423.0 | 1854428.0 | 1902191.0 |
| 2020-06-04 | 1877125.0 | 1902606.0 | 21739.0 | 24296.0 | 1878724.0 | 1926487.0 |
| 2020-06-05 | 1902294.0 | 1926287.0 | 25169.0 | 23682.0 | 1902406.0 | 1950169.0 |
| 2020-06-06 | 1924132.0 | 1948548.0 | 21838.0 | 22261.0 | 1924667.0 | 1972430.0 |
| 2020-06-07 | 1941920.0 | 1969746.0 | 17788.0 | 21198.0 | 1945864.0 | 1993627.0 |
| 2020-06-08 | 1959448.0 | 1990779.0 | 17528.0 | 21033.0 | 1966898.0 | 2014661.0 |
| 2020-06-09 | 1977820.0 | 2012666.0 | 18372.0 | 21887.0 | 1988784.0 | 2036547.0 |
| 2020-06-10 | 1998646.0 | 2035860.0 | 20826.0 | 23194.0 | 2011979.0 | 2059742.0 |
plot_results(us_arima_df, "USA", "incremental ARIMA")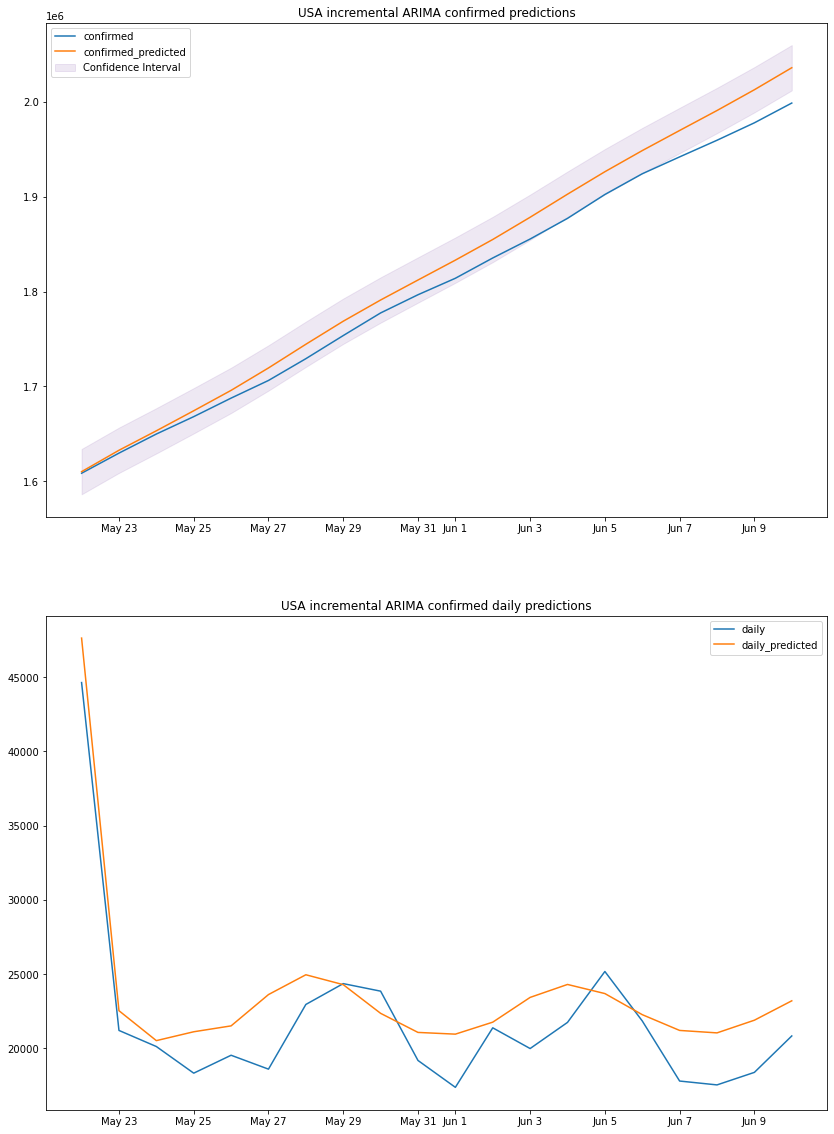
we can see that the predicted totals and predicted daily cases are fairly accurate thus our previous predictions can be taken with some degree of accuracy, and might be used for making decisions.
Conclusion
from all the graphs, functions and numbers above we can come to a simple conclusion that is, there is no single model that will perform best in all scenario even when the data is very similar (in trend not numbers), each model was best for a specific country and wasn’t so far behind in the others for example the HES model is the most accurate with the South Korean dataset but is almost the same as the ARIMA model in Italy.
whats the difference between ARIMA and HES?
ARIMA uses a non-linear function for coefficient calculations, that’s why the
graph does curve sometimes (Italy) while HES is a pure linear method that uses a
linear function and is always a straight line
Considering LSTM is usually the least accurate, is it worth the training
time?
here may be, however, deep learning has its place among machine learning
algorithms and can perform tasks these other functions could never, also the
LSTM model always predicts a wider interval compared to the other 2, in a
practical scenario where range is important the other 2 models will not be ideal
because their results are limited by the original value and don’t spread as
much, the LSTM model could provide better estimates.
ARIMA or HES?
HES, because it takes much less time to train and is as accurate or even more
accurate sometimes.